Polymers · Molecular Dynamics · Damage Mechanics
2025
Self-Healing Polymer Simulator
Watching a material decide to fix itself, atom by atom.
A material that heals itself after damage is not science fiction. It exists. Several classes of polymer networks can close cracks, recover mechanical strength, and repeat the process multiple times over. The question that interested me was not whether they work but why, at the atomic level, and whether you could use that understanding to design better ones without synthesizing a new material every time you wanted to test a hypothesis.
The answer to the second part is yes, but it took months of simulation work to get there.
The two ways a polymer can heal itself
There is an important distinction between extrinsic and intrinsic self-healing, and the distinction matters for what you can simulate.
Extrinsic self-healing uses microcapsules or vascular networks embedded in the material. When a crack forms and propagates, it ruptures the capsules, releasing a liquid healing agent that flows into the crack and polymerizes. The material heals once, or a limited number of times until the local capsule supply is exhausted. The mechanism is macroscopic and relatively well understood. Simulating it requires fluid dynamics and fracture mechanics models, not molecular dynamics.
Intrinsic self-healing is different and more interesting from a molecular design standpoint. Here the healing mechanism is built into the polymer network itself through reversible chemistry. When a crack forms and bonds break, the same functional groups that were covalently linked can reform bonds, either spontaneously through dynamic exchange reactions or when triggered by heat, light, or pH. The material can heal repeatedly, at the same location, as many times as the crack forms. There's no capsule reservoir to exhaust.
The chemistry that enables intrinsic healing typically falls into two categories. The first is reversible covalent bonds: Diels-Alder adducts that dissociate above a threshold temperature and reform on cooling, disulfide bonds that exchange under mild conditions, imine bonds that undergo hydrolysis and reformation. The second is strong non-covalent interactions: quadruple hydrogen bonding motifs like ureidopyrimidinone (UPy) that form such stable complexes they behave almost like covalent bonds at room temperature but can rearrange when the local geometry changes.
I focused on reversible covalent networks, specifically the furan-maleimide Diels-Alder system and disulfide exchange networks, because these have the most direct connection to the vitrimer chemistry I was already modeling for Reformix.
The connection to vitrimer work
This project didn't emerge from nowhere. It grew directly out of the Reformix simulation work. Vitrimers are self-healing materials. Not always by design, but inherently: the same bond exchange mechanism that lets a vitrimer be remolded at elevated temperature also lets it close cracks when the crack faces are brought into contact and heated. The topology rearrangement that enables flow under stress is the same process that enables crack healing.
What I hadn't done for Reformix was simulate the damage event itself: what happens when you apply stress to a crosslinked network, where the bonds break, how the network topology changes as it fails, and whether the healing process actually restores the original network connectivity or just fills the crack with lower-quality structure that will fail again at the same place.
That's what this project addressed.

Setting up the damage simulation
The simulation system is a fully crosslinked polymer network generated using a stepwise crosslinking protocol in LAMMPS. I start with a simulation box of monomer and crosslinker molecules at realistic density, run a short equilibration, then apply a crosslinking algorithm that periodically searches for reactive pairs within a cutoff distance and forms bonds between them, incrementally building up the network. The final system has a crosslink density I can tune by adjusting stoichiometry, and it's structurally realistic in the sense that the network topology reflects actual reaction kinetics rather than an idealized regular lattice.
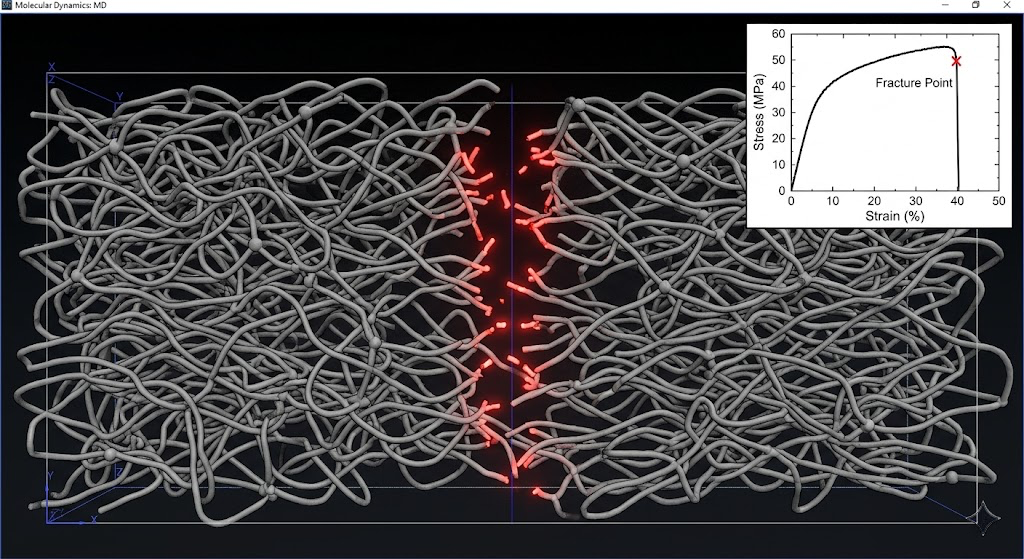
To simulate mechanical damage, I apply a uniaxial tensile deformation by stretching the simulation box at constant strain rate. The stress-strain response shows an initial linear elastic region, yielding, then fracture. For the fracture event to be captured correctly, I need a force field that allows bond breaking. I use a modified Morse potential for the backbone bonds with a dissociation limit, which lets bonds break when they're stretched beyond a threshold. This is a simplification compared to ReaxFF, which models actual reactive chemistry, but it's computationally cheaper and captures the mechanical fracture behavior reasonably well for the purpose of studying network topology changes.
The key output from the damage simulation is the spatial distribution of bond breaks. Where do the bonds fail first? The answer is not random. They fail preferentially at dangling chain ends, at highly strained network junctions where the local topology creates stress concentration, and at the weakest links in the crosslink distribution. A network with a broad distribution of crosslink lengths, where some chains are much shorter than others, fails at the short chains first because they bear disproportionate stress. This is why polydispersity in chain length between crosslinks degrades fracture toughness.
Simulating the healing event
After the damage simulation, I have a cracked network. To simulate healing, I bring the two crack faces back together by reversing the deformation, then raise the temperature above the threshold for bond exchange. In the furan-maleimide system, this is around 120°C. Above this temperature, the retro-Diels-Alder reaction breaks the adducts, generating free furan and maleimide groups. These can diffuse, find new partners, and reform adducts in a different configuration.
The diffusion part is the challenge. In a dense crosslinked network, chain mobility is severely restricted. A free furan group at a crack surface can only sample configurations within a limited range determined by the length of the chain it's attached to and the local network topology. Whether it finds a maleimide partner depends on whether there's one within reach. This is fundamentally a topological problem: the healing efficiency is controlled not just by the reactivity of the functional groups but by the local connectivity of the network.
I quantify this through a bond-reformation tracking algorithm. After each healing simulation timestep, I check whether any broken bonds have reformed and record the new bond topology. At the end of the healing simulation, I compute two numbers. The first is the chemical healing efficiency: what fraction of broken bonds reformed. The second is the mechanical healing efficiency: what is the Young's modulus of the healed network relative to the pristine network. These two numbers are not the same. You can reform 90% of the broken bonds and recover only 60% of the stiffness if the reformed bonds happen to be in geometrically unfavorable configurations that don't carry load effectively.
The gap between chemical and mechanical healing efficiency is one of the most practically important outputs of the simulation. It tells you whether a material is limited by reaction kinetics (can't reform bonds fast enough) or by network topology (reforms bonds but in the wrong places). These failure modes have different solutions. If it's kinetics, you need a faster exchange chemistry or higher temperature. If it's topology, you need a different crosslink architecture.
What the simulations showed
The furan-maleimide system heals well in simulation under ideal conditions, meaning the crack faces are in direct contact, there's no contamination, and the temperature is well above the retro-DA threshold. Chemical healing efficiency reaches 80 to 85% after two hours at 130°C in the simulation. Mechanical recovery is lower, typically 65 to 75% of original modulus, confirming the topological gap.
The most important variable I found is the crosslink density. Very highly crosslinked networks fracture cleanly but heal poorly because the chain segments are too short to allow the end groups to find new partners. Very lightly crosslinked networks heal well but have poor initial mechanical properties. There's an optimum in the middle where both properties are acceptable, and the simulation locates it more precisely than intuition alone would. For the furan-maleimide chemistry with the chain lengths I simulated, the optimum crosslink density was around 15 to 20% of available sites, which corresponds to a gel fraction above 95% and moderate rubbery modulus.
Disulfide exchange networks show a different character. The exchange chemistry is faster and lower temperature (around 60°C is sufficient for many disulfide-based systems), which means healing doesn't require the material to go above its glass transition temperature. You can heal at temperatures where the network is still glassy, which is important for applications where maintaining structural rigidity during healing matters. The tradeoff is that disulfide bonds are weaker than carbon-carbon backbone bonds, which limits how much of the original strength can be recovered. My simulations confirmed this: the disulfide networks healed faster and at lower temperature but recovered only about 55% of the original modulus under the same conditions where the furan-maleimide system recovered 70%.
The simulation as design tool
The practical use of this project is not to explain why existing materials heal. It's to test design modifications before synthesizing them.
I ran a series of virtual experiments: what happens if you increase the furan functionality from two to three per chain? What if you add a small amount of non-reactive diluent to increase local chain mobility near the crack surface? What if you use a bimodal crosslink length distribution instead of a narrow one? Each of these is a parameter change in the simulation that maps to a concrete synthetic decision.
The trimerization result was the most useful: going from difunctional to trifunctional furan groups at the chain ends improved mechanical healing efficiency from 72% to 81% in simulation, because the higher functionality means each chain end has more potential partners within its reachable volume. This is a testable prediction. The synthesis of trifunctional furan-terminated chains is not trivial but it's not unknown. Someone with a synthesis lab could test it directly, and the simulation gives them a specific prediction for what the healing efficiency improvement should be.
That's what I want computational work in materials to do. Not just explain the past but generate specific, falsifiable predictions about materials that don't exist yet. Self-healing polymer design has historically been driven by chemical intuition and trial-and-error synthesis screening. Simulation doesn't replace that, but it can front-load some of the iteration into compute time, which is cheaper than lab time. The predictions sit in a document. The experiments that will confirm or refute them are the next step.